 |
The Sullivan Lab, MCD Biology, UCSC |

| ||
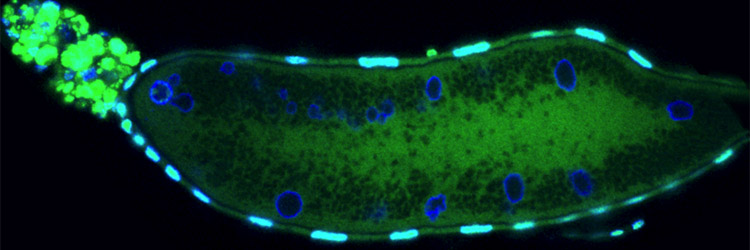
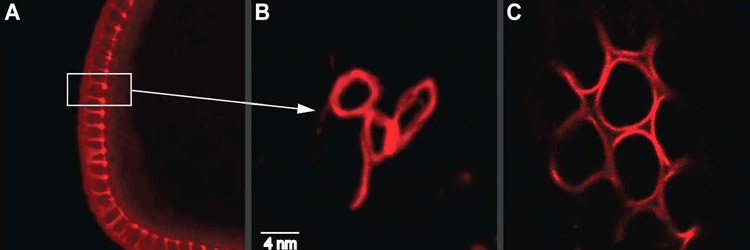
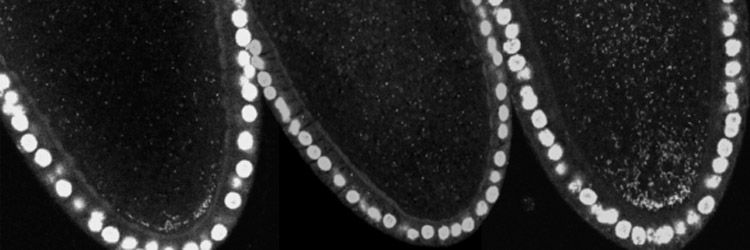
Sullivan Lab Research IV. Revisiting Past Projects Below is a list of projects that our lab has worked and published on. We continue to have a great interest in these biological issues and will likely revisit them when the opportunities arises. Cell Biology of Parthenogenesis: Parthenogenesis refers to the ability of embryos to develop without being fertilized. Parthenogenesis is typically a derived trait and has independently evolved numerous times from non-parthenogenetic ancestors. For example it is estimated that parthenogenetic haplo-dipoid insects have independently evolved at least 17 times. In another striking example, the New Zealand water snail has independently evolved parthenogenesis over 100 times in lakes less than 100,000 years old. These findings are astonishing when one considers the dramatic alterations that must occur at the cellular level for an egg to successfully develop in the absence of a sperm. The sperm provides the paternal chromosome complement and a basal body, which immediately transforms into a centrosome upon fertilization. Therefore evolution of parthenogenesis required an ability to compensate for the lack of a paternal genome and an ability to generate centrosomes solely from maternally supplied products. We discovered that hundreds of centrosomes form de novo in specialized lamin-based organelles derived from the nucleus of the female oocyte. The Hunt for Natural Genetic Wolbachia Variants Wolbachia—an obligate, intracellular bacteria—infect millions of insect and nematode hosts, impacting human health worldwide. For example, the pathogenic nematodes that cause elephantiasis and onchocerciases (river blindness) require Wolbachia as mutualist endosymbionts, and Wolbachia contribute to the pathogenesis of these disease. Furthermore, an ongoing, novel vector control strategy uses Wolbachia-infected mosquitoes to prevent transmission of the tropical disease dengue fever. Ecologists, evolutionary biologists, and population geneticists have extensively studied Wolbachia. Recently, researchers began investigating the molecular and cellular biology of Wolbachia-host interactions, including maternal transmission of Wolbachia, factors controlling Wolbachia titer and tissue distribution, and cell-to-cell invasion by Wolbachia. Using the powerful genetic tools available in Drosophila, our lab and others have begun to dissect the host side of these interactions. However, the thus far genetic intractability of Wolbachia has prevented investigating the endosymbiont side of these interactions. In fact, the lack of Wolbachia mutants or variants is a primary limitation to further characterizing the molecular and cellular basis of Wolbachia-host interactions. One solution is to identify natural Wolbachia variants by collecting and screening wild-populations of Drosophila. To this end we have surveyed wild populations in Southern Big Sur and the Channel Islands. These studies have yielded variants in Wolbachia morphology, tissue distribution and titer. Unfortunately most of these variants have proven unstable and after maintenance in the lab for over a year they are no longer present. Are recent efforts in collaboration with the Tureelli lab at UC Davis suggest that Wolbachia variants in distribution and titer in the adult brain are much more stable. Testing the Behavior Modification Hypothesis: Our studies demonstrate that a conserved feature not only is localization to the germline but also the adult central brain. In addition, different strains of Wolbachia target different regions of the central brain. This suggests specific targeting mechanisms are involved. In fact we find Wolbachia exhibits a dramatic asymmetric division in order to maintain as association with the larval neuroblast stem cell ensuring its incorporation into the adult nervous system. The functional significance of this neuronal localization remains unclear. One intriguing possibility is that Wolbachia may alter behavior to promote its own transmission (in host/symbiont literature this is known as the behavioral modification hypothesis). Support for this notion is that Wolbachia infected females are more promicious than uninfected females and even readily mate with males from different species. One means of determining whether this effect is a direct result of Wolbachia localization in the brain in examine different strains of Wolbachia with different pattersn of brain localization and determine whether the behavioral effects on Drosophila are correlated with localization. Cellular Constraints and Adaptation to Changes in Karyoptypens Why do butterflies have 100s of chromosomes and nematodes such as ascaris only one? Does the size and shape of the spindle and cell correlate with karyotype? Can the cell and spindle adapt to the dramatic alterations in karyotype brought on by large-scale chromosome rearrangements? We have begun to address these questions by taking advantage of a vast well characterized array of Drosophila lines bearing translocations, inversions and chromosome rearrangements. We have focused on studies on a compound chromosomes in which both second chromosomes are fused to a single centromere. This results in chromosomes with arms twice the normal length. Live analysis reveals that the cells and spindle elongate in order to provide sufficient space to segregate the longer arms of the compound chromosome. Representative Publications in these Areas Ferree PM, McDonald K, Fasulo B, Sullivan WT. (2006) The origin of Centrosomes in Parthenogenetic Hymenopteran Insects. Current Biol. 16: 801-807.[PDF] Tram, U. and W. Sullivan. (2000) Reciprocal inheritance of centrosomes in the parthenogenetic Hymenopteran Nasonia vitripennis. Current Biology10:1413-1419. [PDF] Casper-Lindley C, Kimura S, Saxton DS, Essaw Y, Simpson I, Tan V, Sullivan W. (2011) Fluorescent-based method for rapid Wolbachia detection in the Drosophila germline and somatic tissues. Appl Environ Microbiol. 17 (44): 4788-4794. [PDF] For a complete list of Sullivan lab publications, see our main Publications page [Back] [Next] [Sullivan Research Main Page ] |
||
|
|